 The goal is to find homologous sequences from which properties of the query sequence can be inferred, such as molecular and cellular functions and structure. The most widely used approach to protein annotation and analysis is based on sequence similarity search 5, 6, 7, 8. The scale of these databases poses challenges to state-of-the-art analysis methods. 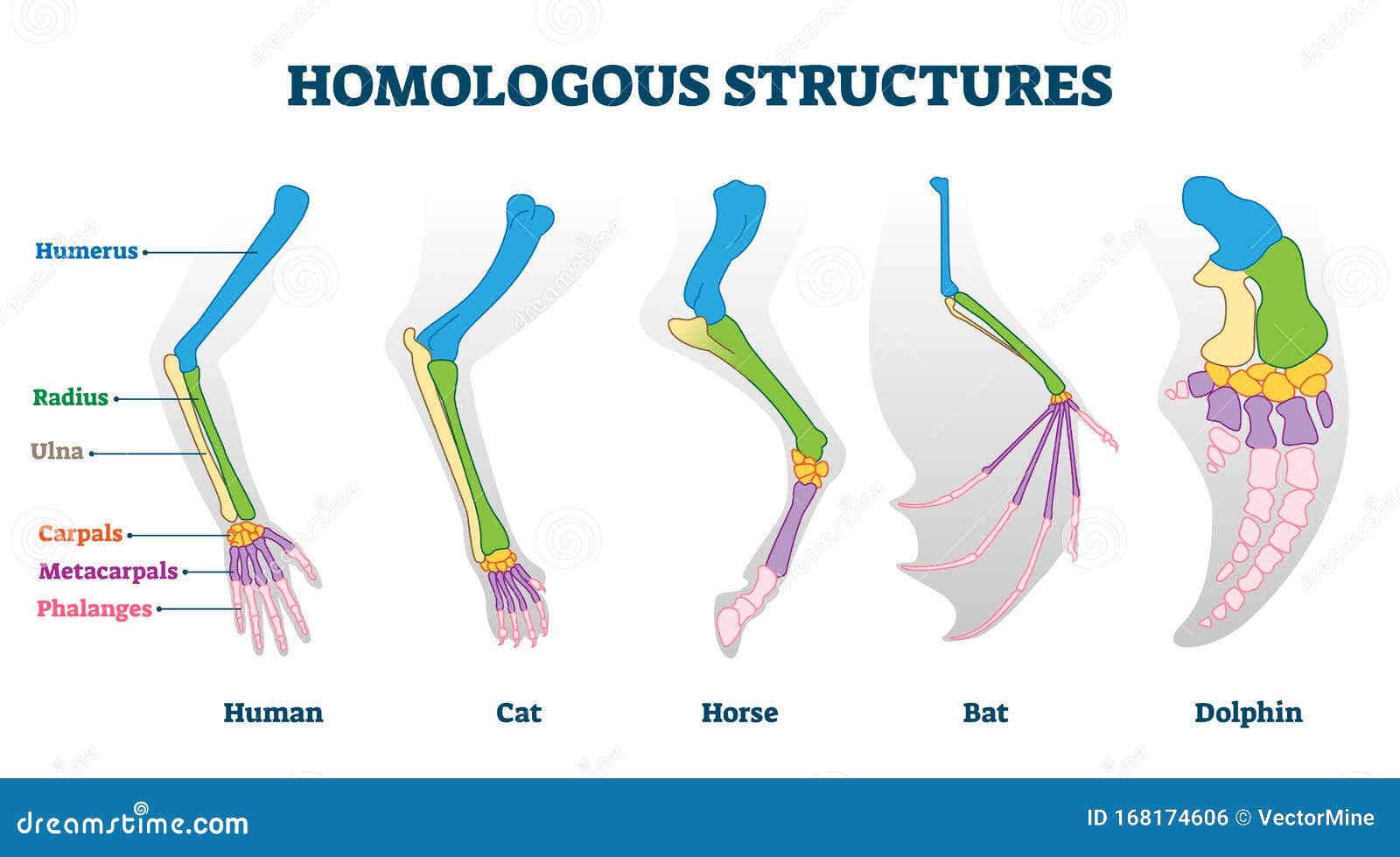
3), and the ESM Atlas contains over 617 million metagenomic structures predicted by ESMFold 4. The European Bioinformatics Institute already holds over 214 million structures predicted by AlphaFold2 (ref.  
The recent developments in in silico protein structure prediction at near-experimental quality 1, 2 are advancing structural biology and bioinformatics. 
0 Comments
Leave a Reply. |

 RSS Feed
RSS Feed
